Quantitative analysis of tRNA abundance and modifications by nanopore RNA sequencing
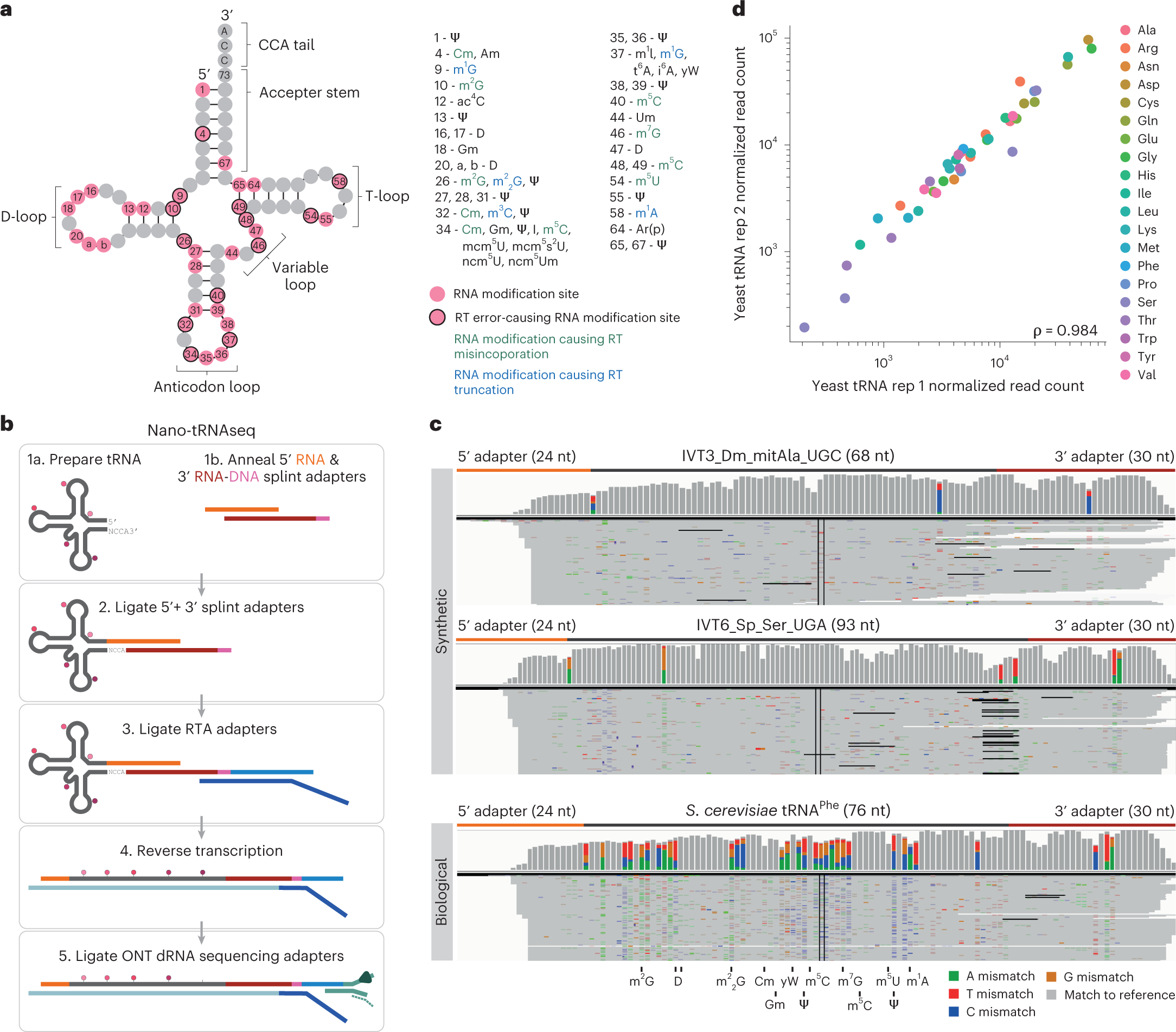

Combining tRNA sequencing methods to characterize plant tRNA expression and post-transcriptional modification
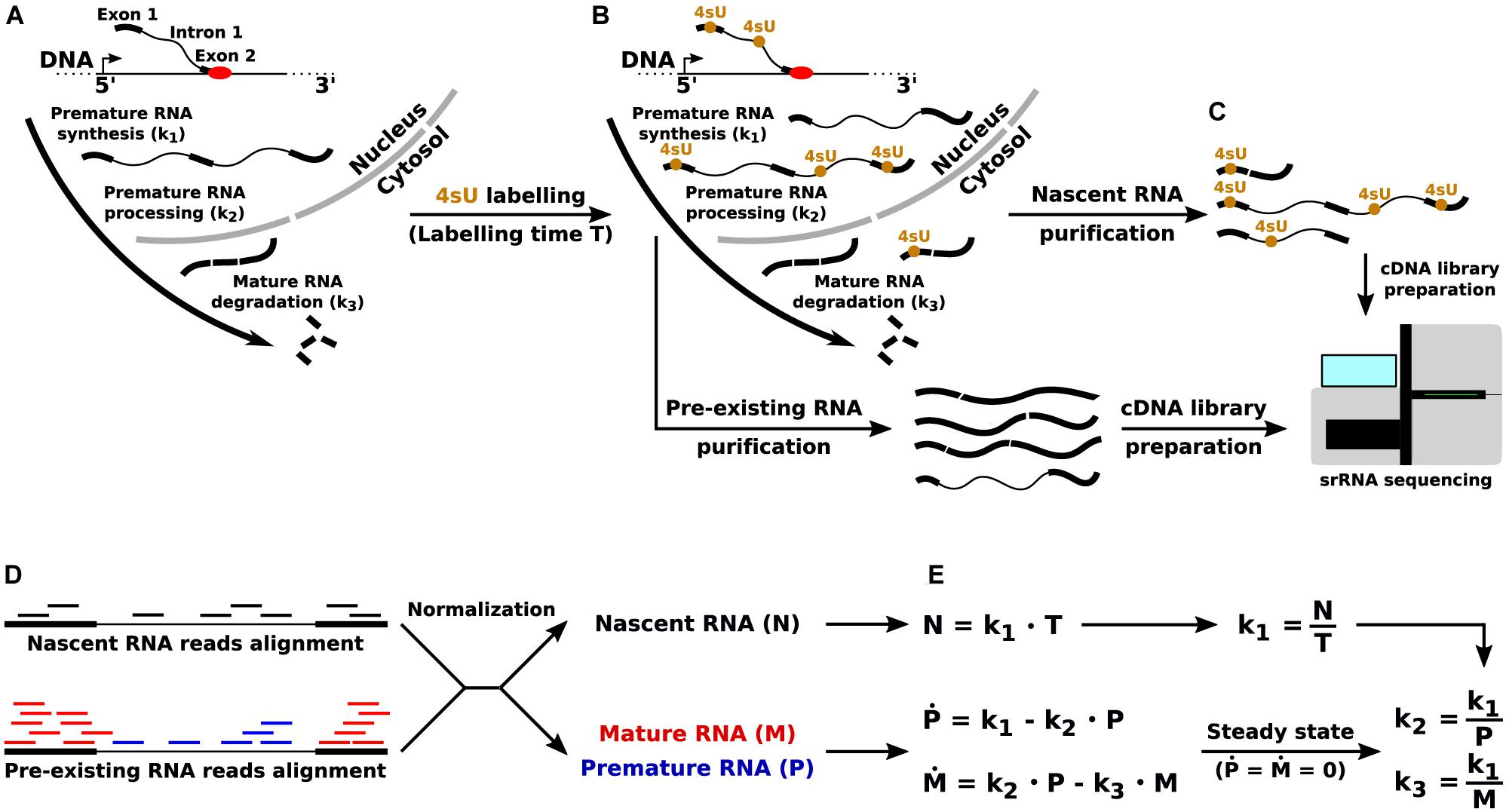
Frontiers Direct RNA Sequencing for the Study of Synthesis, Processing, and Degradation of Modified Transcripts
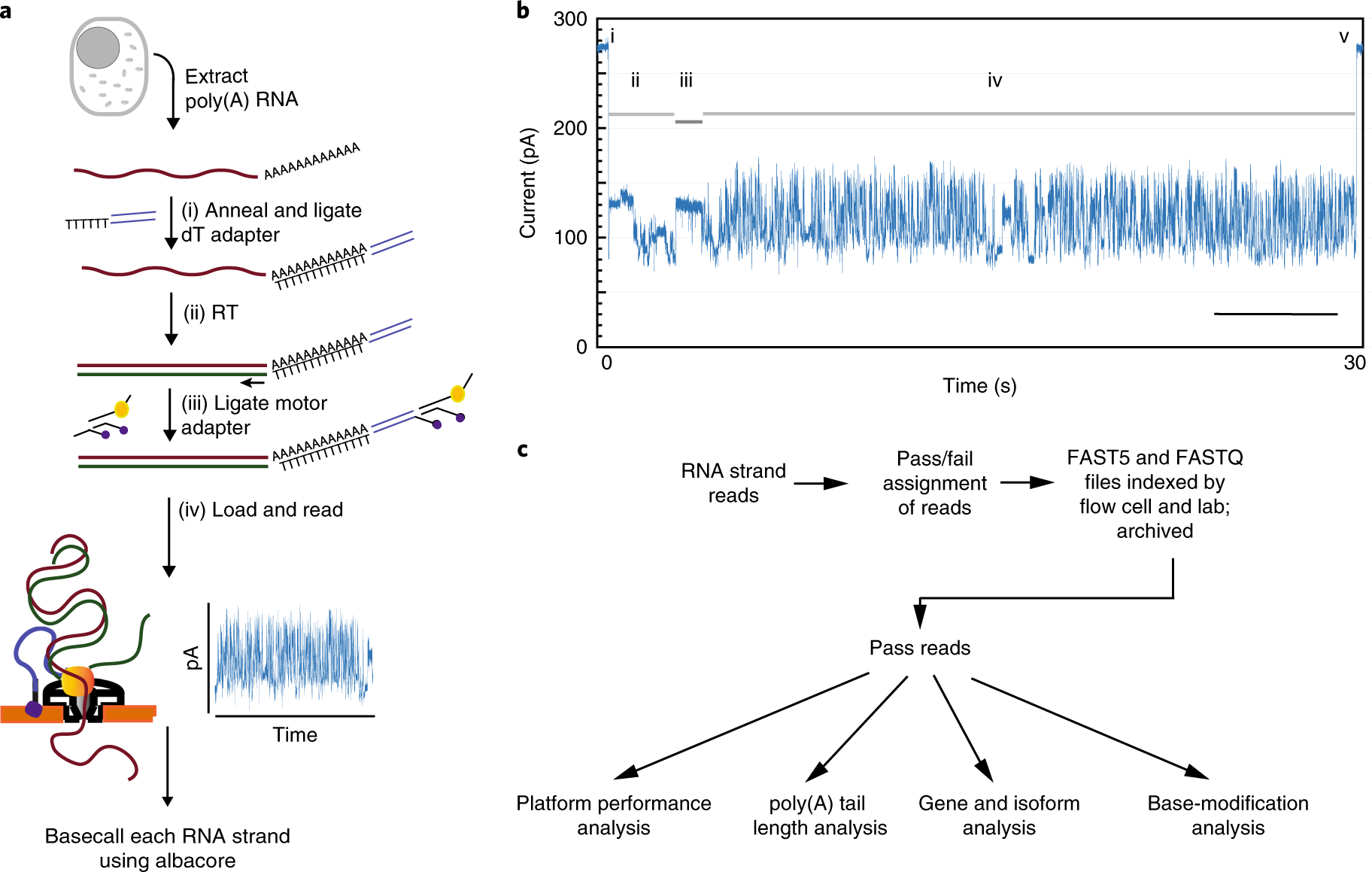
Nanopore native RNA sequencing of a human poly(A) transcriptome

Expanding the epitranscriptomic RNA sequencing and modification mapping mass spectrometry toolbox with field asymmetric waveform ion mobility and electrochemical elution liquid chromatography

Quantitative analysis of tRNA abundance and modifications by nanopore RNA sequencing

Overview of AQRNA-seq a, Library preparation workflow. b–d
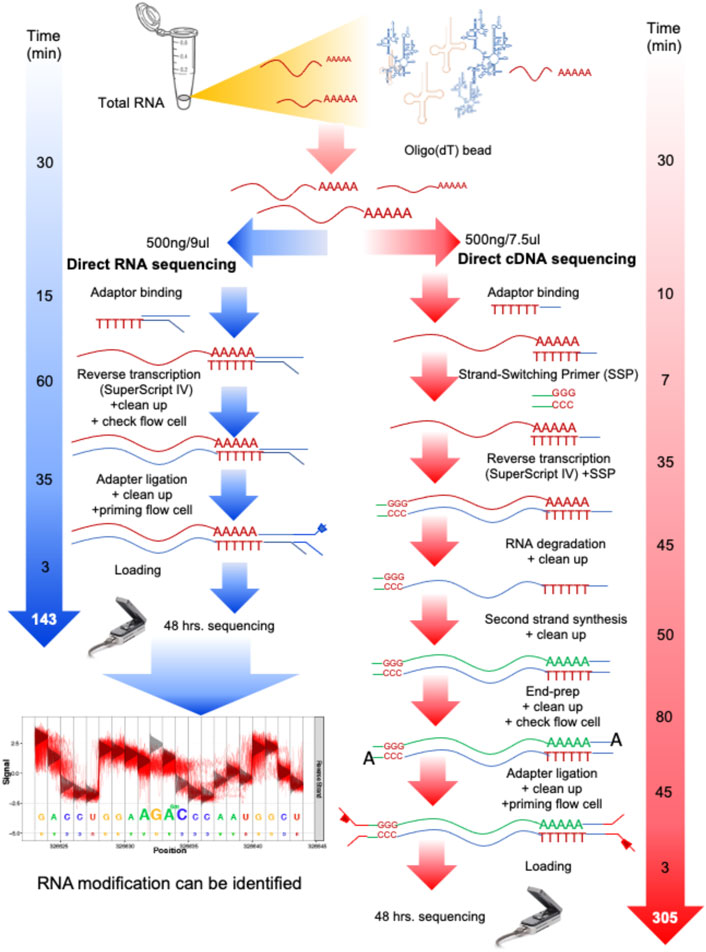
Frontiers Native RNA or cDNA Sequencing for Transcriptomic Analysis: A Case Study on Saccharomyces cerevisiae
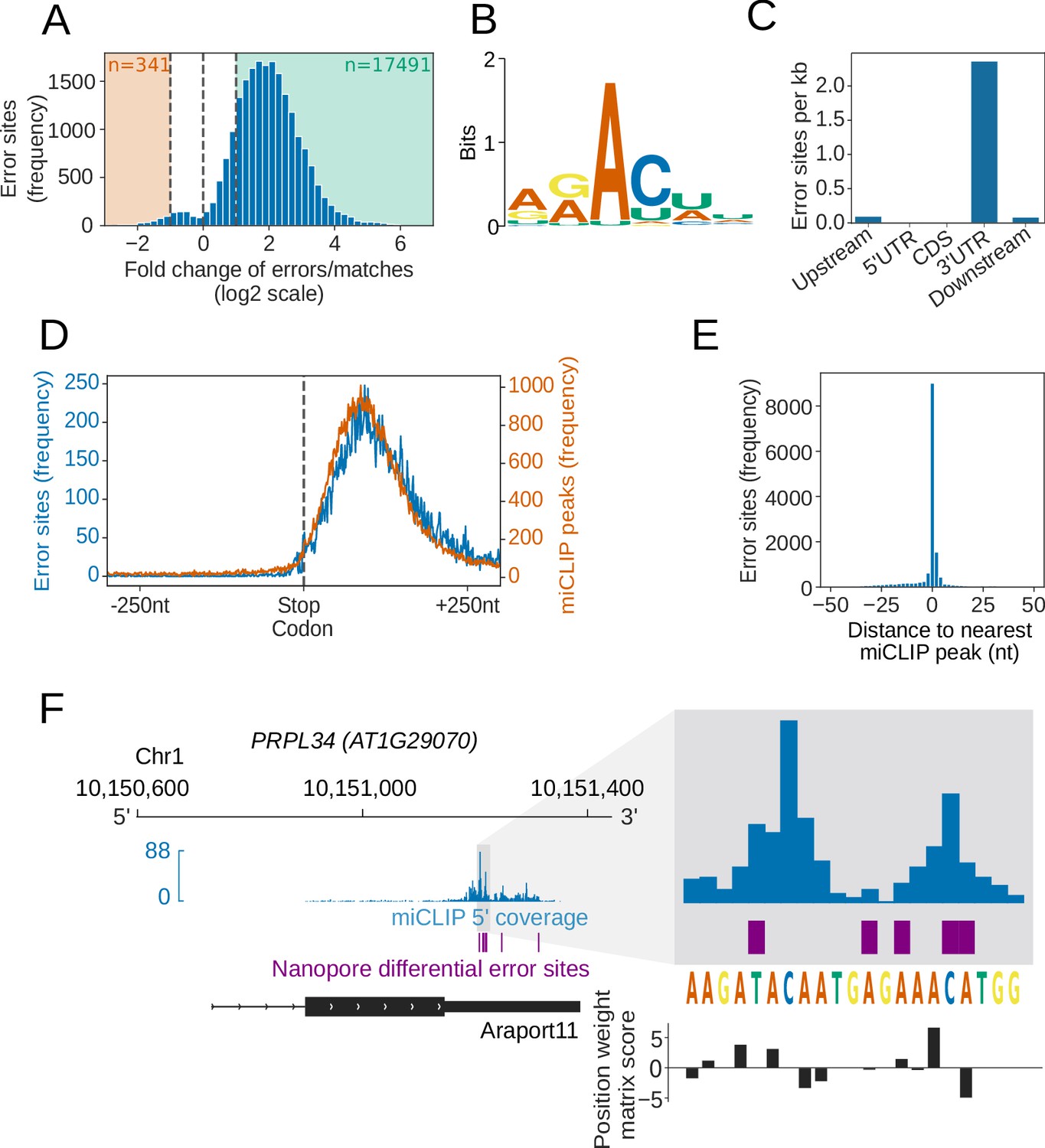
Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification

Full article: Combining tRNA sequencing methods to characterize plant tRNA expression and post-transcriptional modification

Quantitative profiling of pseudouridylation dynamics in native RNAs with nanopore sequencing

Full article: Combining tRNA sequencing methods to characterize plant tRNA expression and post-transcriptional modification

Direct Nanopore Sequencing of Individual Full Length tRNA Strands

Quantitative analysis of tRNA abundance and modifications by nanopore RNA sequencing

Quantitative tRNA-sequencing uncovers metazoan tissue-specific tRNA regulation

Nanopore-based native RNA sequencing provides insights into prokaryotic transcription, operon structures, rRNA maturation and modifications









